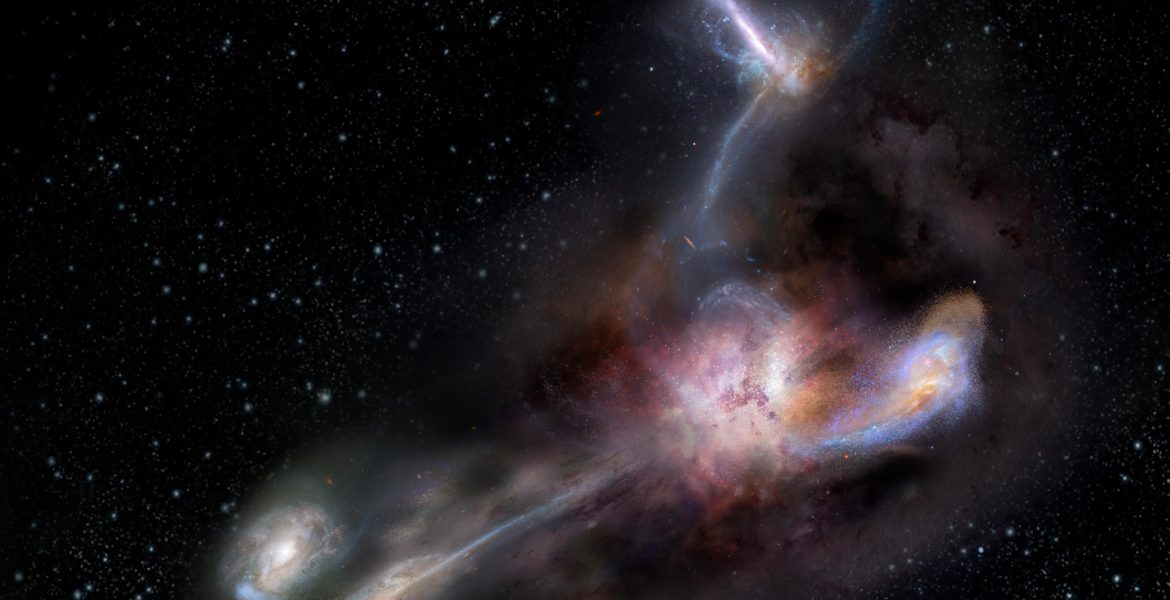
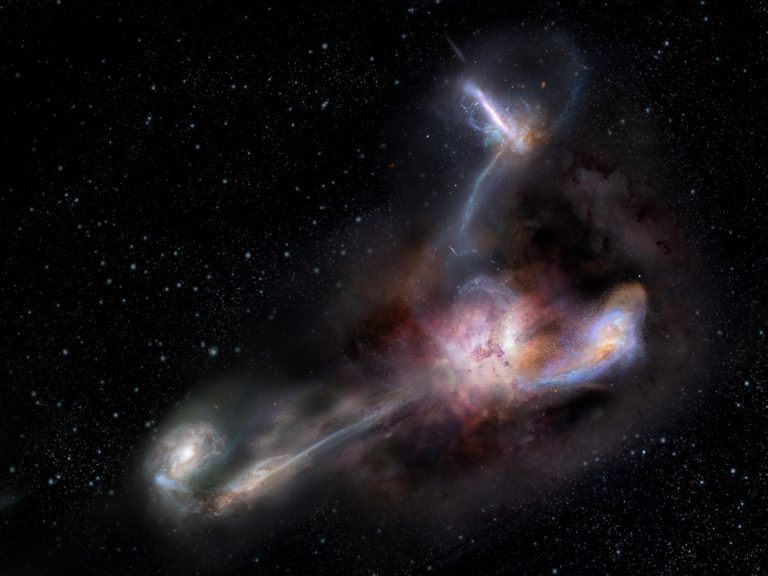
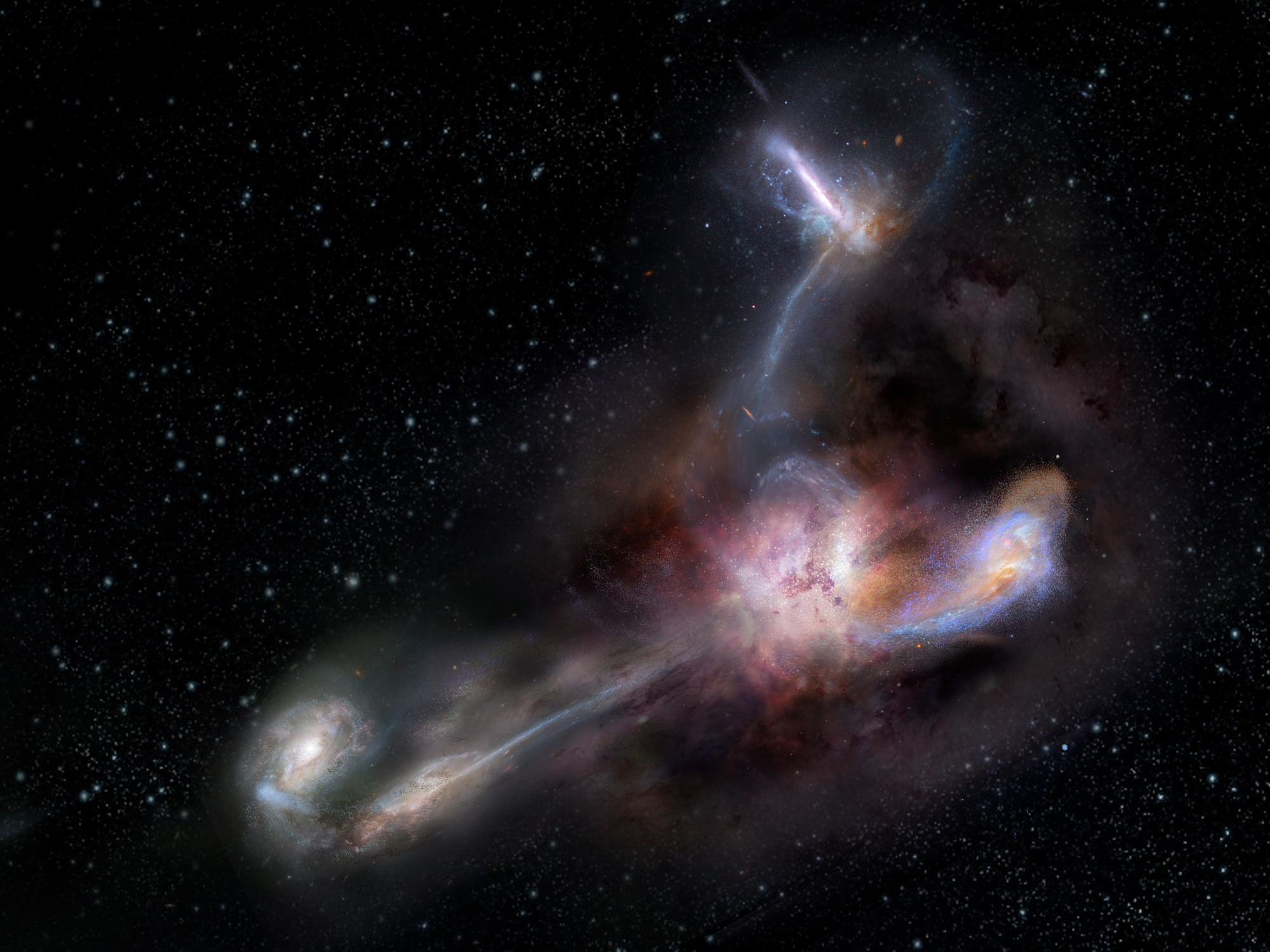
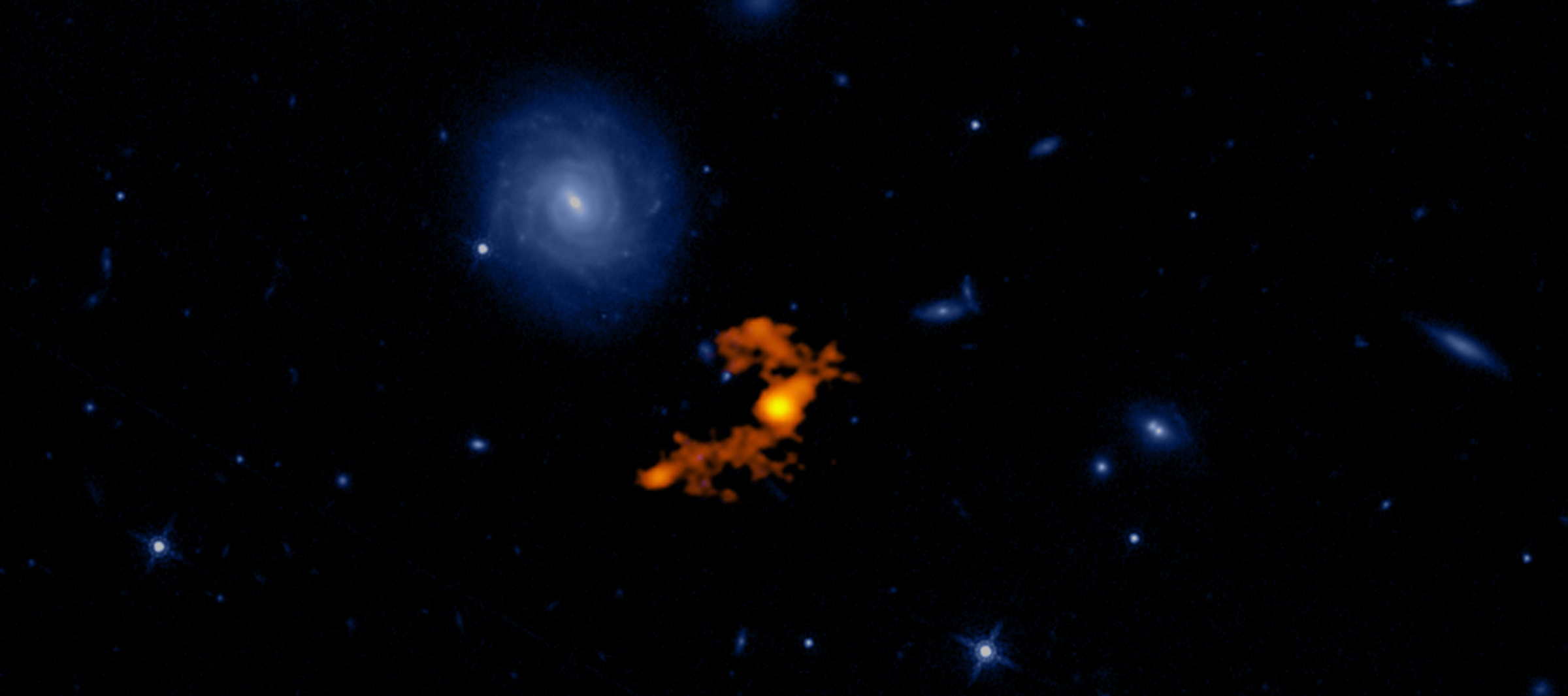
The most luminousLuminousIn astronomy, luminosity is an actual unit. It's the amount of energy given off by a star or galaxy, or any other object -- over a specific period of time. "Brightness," in astronomy, is how apparently bright an object appears to an observer. Brightness goes down with distance. Luminosity does not. galaxy in the universe has been caught in the act of stripping away nearly half the mass from at least three of its smaller neighbors, according to a new study published in the journal Science. The light from this galaxy, known as W2246-0526, took 12.4 billion years to reach us, so we are seeing it as it was when our universe was only about a tenth of its present age.
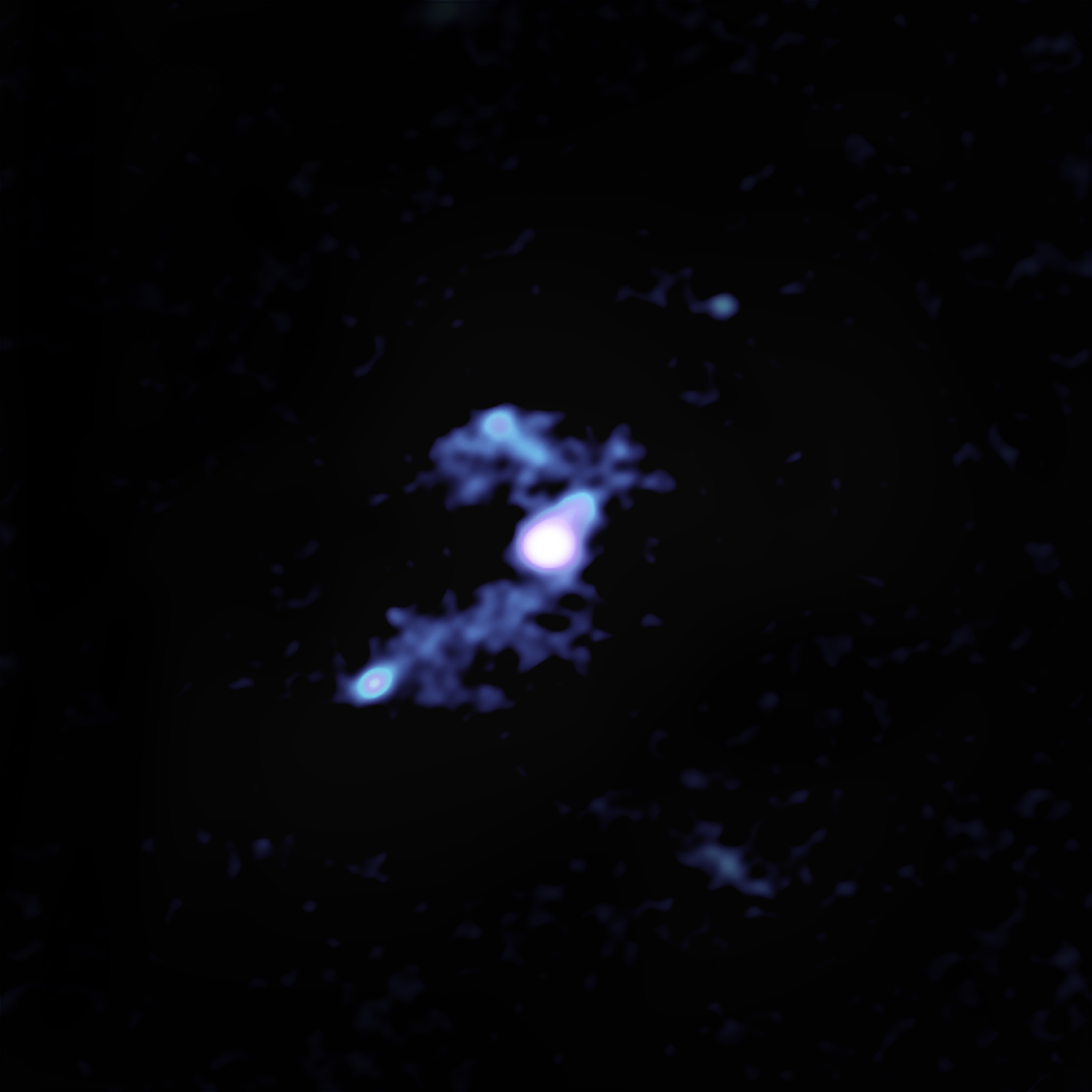
New observations with the Atacama Large Millimeter/submillimeter Array (ALMA)Atacama Large Millimeter/submillimeter Array (ALMA)Funded by the U.S. National Science Foundation and its international partners (NRAO/ESO/NAOJ), ALMA is among the most complex and powerful astronomical observatories on Earth or in space. The telescope is an array of 66 high-precision dish antennas in northern Chile. reveal distinct streamers of material being pulled from three smaller galaxies and flowing into the more massive galaxy, which was discovered in 2015 by NASA’s space-based Wide-field Infrared Survey Explorer (WISE). It is by no means the largest or most massive galaxy we know of, but it is unrivaled in its brightness, emitting as much infrared light as 350 trillion Suns.
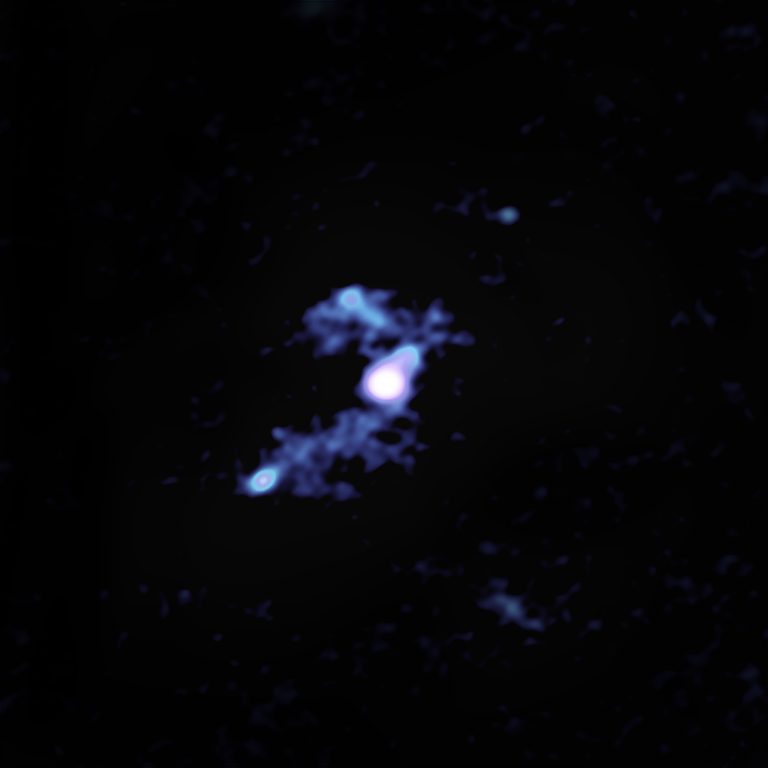
The connecting tendrils between the galaxies contain about as much material as the galaxies themselves. ALMA’s amazing resolution and sensitivity allowed the researchers to detect these remarkably faint and distant trans-galactic streamers.
“We knew from previous data that there were three companion galaxies, but there was no evidence of interactions between these neighbors and the central source,” said Tanio Díaz-Santos of the Universidad Diego Portales in Santiago, Chile, lead author of the study. “We weren’t looking for cannibalistic behavior and weren’t expecting it, but this deep dive with the ALMA observatory makes it very clear.”
Galactic cannibalism is not uncommon, though this is the most distant galaxy in which such behavior has been observed and the study authors are not aware of any other direct images of a galaxy simultaneously feeding on material from multiple sources at those early cosmic times.
The researchers emphasize that the amount of gas being devoured by W2246-0526 is enough to keep it forming stars and feeding its central black hole for hundreds of millions of years.
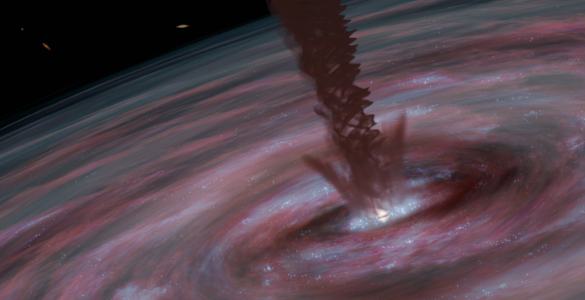
This galaxy’s startling luminosity is not due to its individual stars. Rather, its brightness is powered by a tiny, yet fantastically energetic disk of gas that is being superheated as it spirals in on the supermassive black holeSupermassive Black HoleA black hole that has a million or as much as a billion solar masses. These large black holes lurk at the centers of most galaxies.. The light from this blazingly bright accretion diskAccretion diskA disk of gas that accumulates around a center of gravitational attraction, such as a white dwarf, a neutron star, or a black hole. As the gas spirals in, it becomes hot and emits energy at a variety of wavelengths, including X-ray and radio waves. is then absorbed by the surrounding dust, which re-emits the energy as infrared light.
This extreme infrared radiation makes this galaxy one of a rare class of quasarsQuasarAn apparently small (at least to observers on Earth) yet immensely powerful cosmic object. Some quasars (quasi-stellar objects, or QSOs) are strong radio sources. Radio-emitting quasars were the first to be discovered. These are some of the most distant objects in the Universe, and are believed to be fueled by supermassive black holes residing in ancient galaxies. known as Hot, Dust-Obscured Galaxies or Hot DOGs. Only about one out of every 3,000 quasarsQuasarAn apparently small (at least to observers on Earth) yet immensely powerful cosmic object. Some quasars (quasi-stellar objects, or QSOs) are strong radio sources. Radio-emitting quasars were the first to be discovered. These are some of the most distant objects in the Universe, and are believed to be fueled by supermassive black holes residing in ancient galaxies. observed by WISE belongs to this class.
Much of the dust and gas being siphoned away from the three smaller galaxies is likely being converted into new stars and feeding the larger galaxy’s central black hole. This galaxy’s gluttony, however, may lead to its self-destruction. Previous research suggests that the energy of the AGN will ultimately jettison much, if not all of the galaxy’s star-forming fuel.
An earlier work led by co-author Chao-Wei Tsai of UCLA estimates that the black hole at the center of W2246-0526 is about 4 billion times the mass of the Sun. The mass of the black hole directly influences how bright the AGN can become, but — according to this earlier research — W2246-0526 is about 3 times more luminous than what should be possible. Solving this apparent contradiction will require additional observations.
The National Radio Astronomy Observatory is a facility of the National Science Foundation, operated under cooperative agreement by Associated Universities, Inc.
# # #
Contact:
Charles Blue, Public Information Officer
(434) 296-0314; cblue@nrao.edu
This research is presented in a paper titled “The Multiple Merger Assembly of a Hyper-luminous Obscured Quasar at redshift 4.6 ” by T. Díaz-Santos et al. in the journal Science. [http://science.sciencemag.org/cgi/doi/10.1126/science.aap7605]
The Atacama Large Millimeter/submillimeter Array (ALMA), an international astronomy facility, is a partnership of the European Southern Observatory (ESO), the U.S. National Science Foundation (NSF) and the National Institutes of Natural Sciences (NINS) of Japan in cooperation with the Republic of Chile. ALMA is funded by ESO on behalf of its Member States, by NSF in cooperation with the National Research Council of Canada (NRC) and the Ministry of Science and Technology (MOST) in Taiwan and by NINS in cooperation with the Academia Sinica (AS) in Taiwan and the Korea Astronomy and Space Science Institute (KASI).
ALMA construction and operations are led by ESO on behalf of its Member States; by the National Radio Astronomy Observatory (NRAO), managed by Associated Universities, Inc. (AUI), on behalf of North America; and by the National Astronomical Observatory of Japan (NAOJ) on behalf of East Asia. The Joint ALMA Observatory (JAO) provides the unified leadership and management of the construction, commissioning and operation of ALMA.